DREAMS: deep read-level error model for sequencing data applied to low-frequency variant calling and circulating tumor DNA detection, Genome Biology
Por um escritor misterioso
Last updated 12 abril 2025
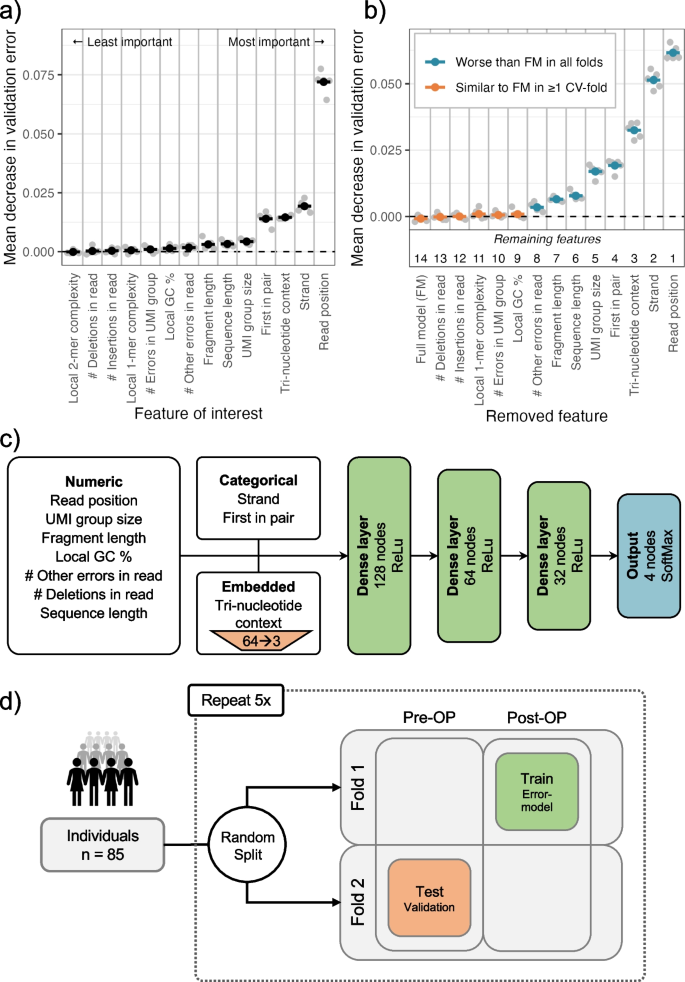
Circulating tumor DNA detection using next-generation sequencing (NGS) data of plasma DNA is promising for cancer identification and characterization. However, the tumor signal in the blood is often low and difficult to distinguish from errors. We present DREAMS (Deep Read-level Modelling of Sequencing-errors) for estimating error rates of individual read positions. Using DREAMS, we develop statistical methods for variant calling (DREAMS-vc) and cancer detection (DREAMS-cc). For evaluation, we generate deep targeted NGS data of matching tumor and plasma DNA from 85 colorectal cancer patients. The DREAMS approach performs better than state-of-the-art methods for variant calling and cancer detection.

LFMD: detecting low-frequency mutations in high-depth genome
Frontiers Benchmarking Low-Frequency Variant Calling With Long

DREAMS: Deep Read-level Error Model for Sequencing data applied to

Detecting and Quantitating Low Fraction DNA Variants with Low
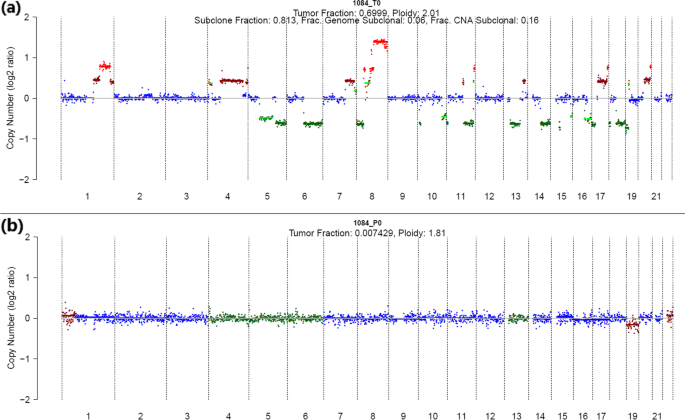
Whole genome deep sequencing analysis of cell-free DNA in samples
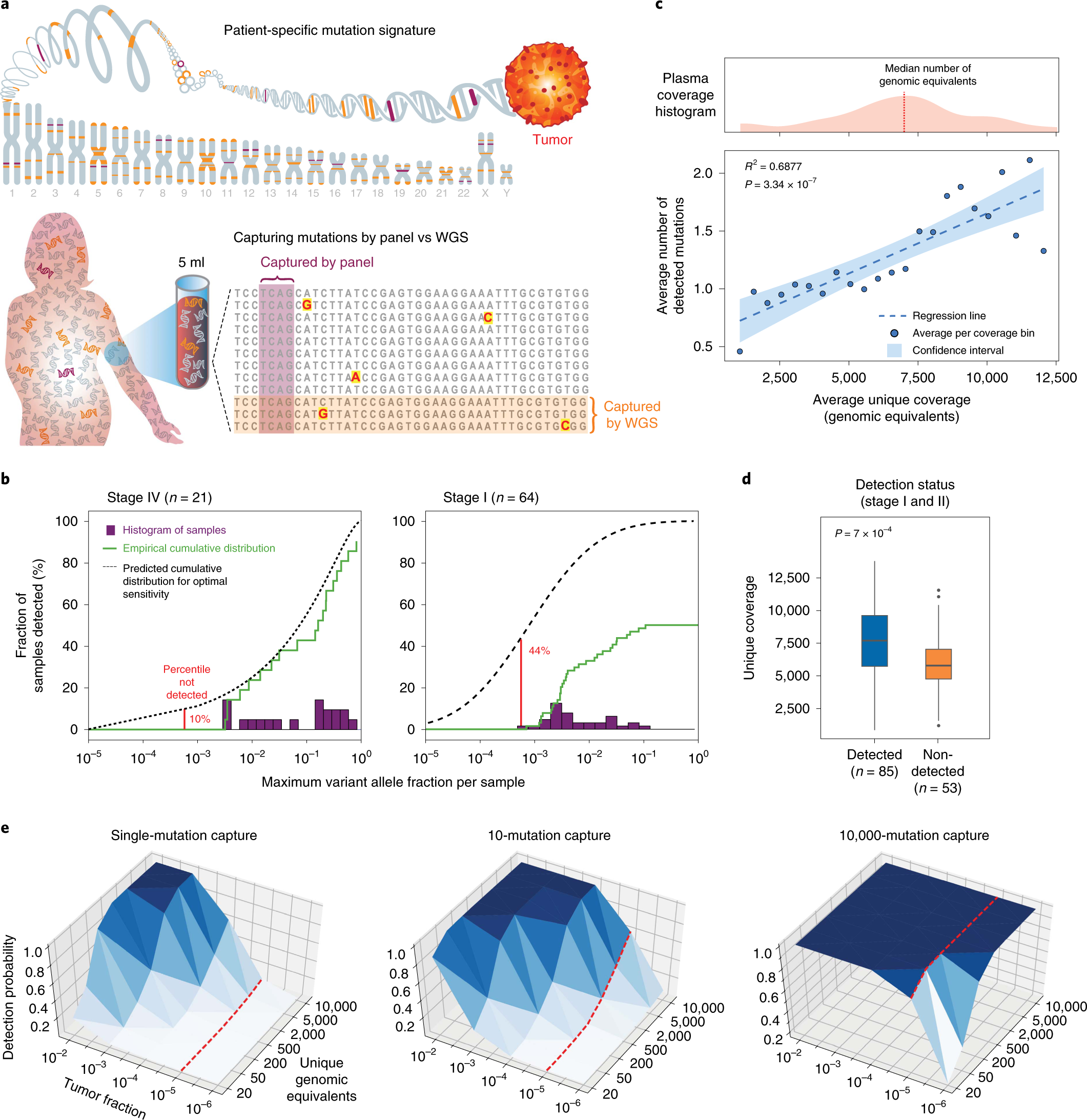
Genome-wide cell-free DNA mutational integration enables ultra

DREAMS: deep read-level error model for sequencing data applied to

The changing face of circulating tumor DNA (ctDNA) profiling

Applications and analysis of targeted genomic sequencing in cancer
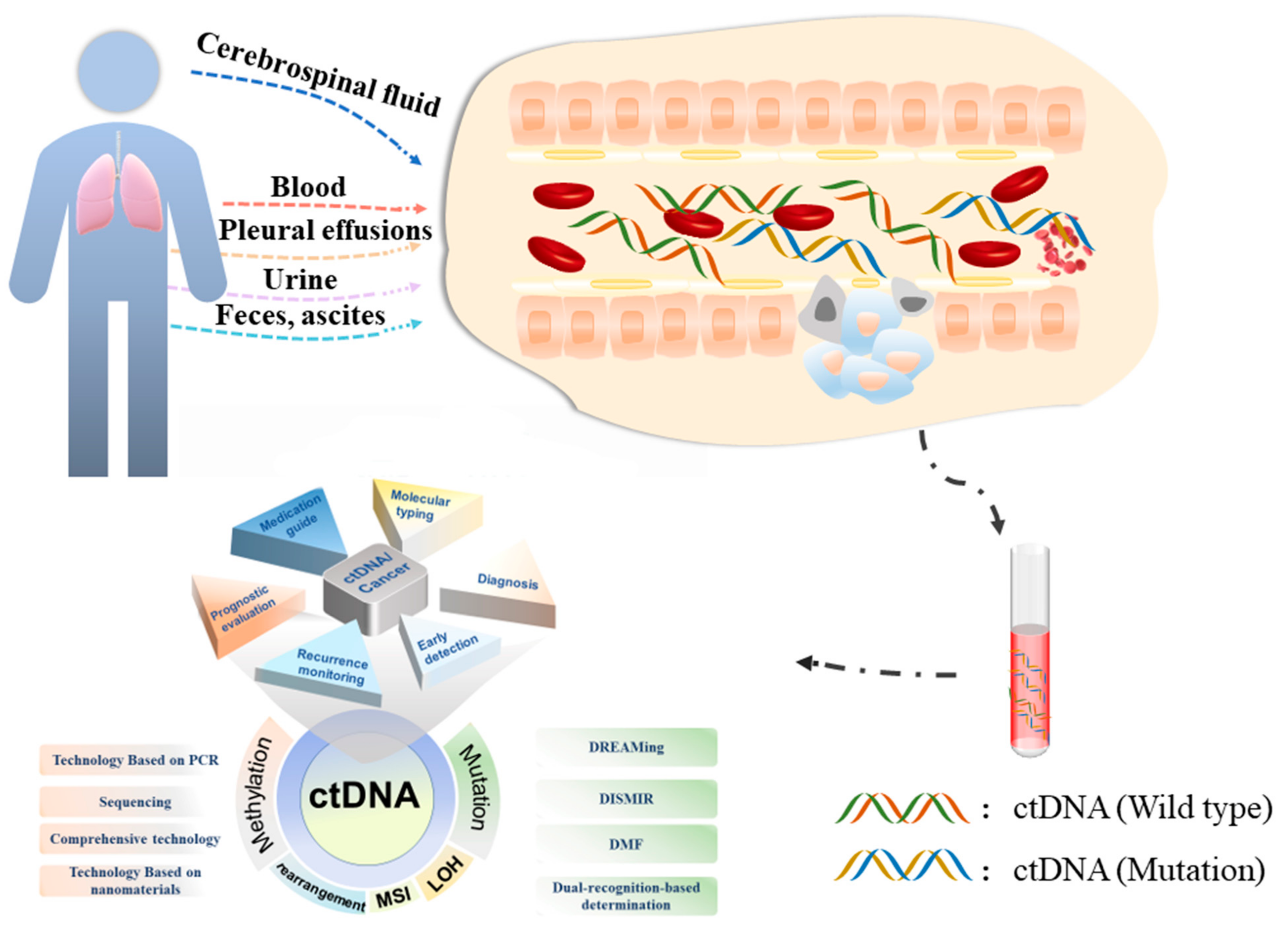
Cancers, Free Full-Text
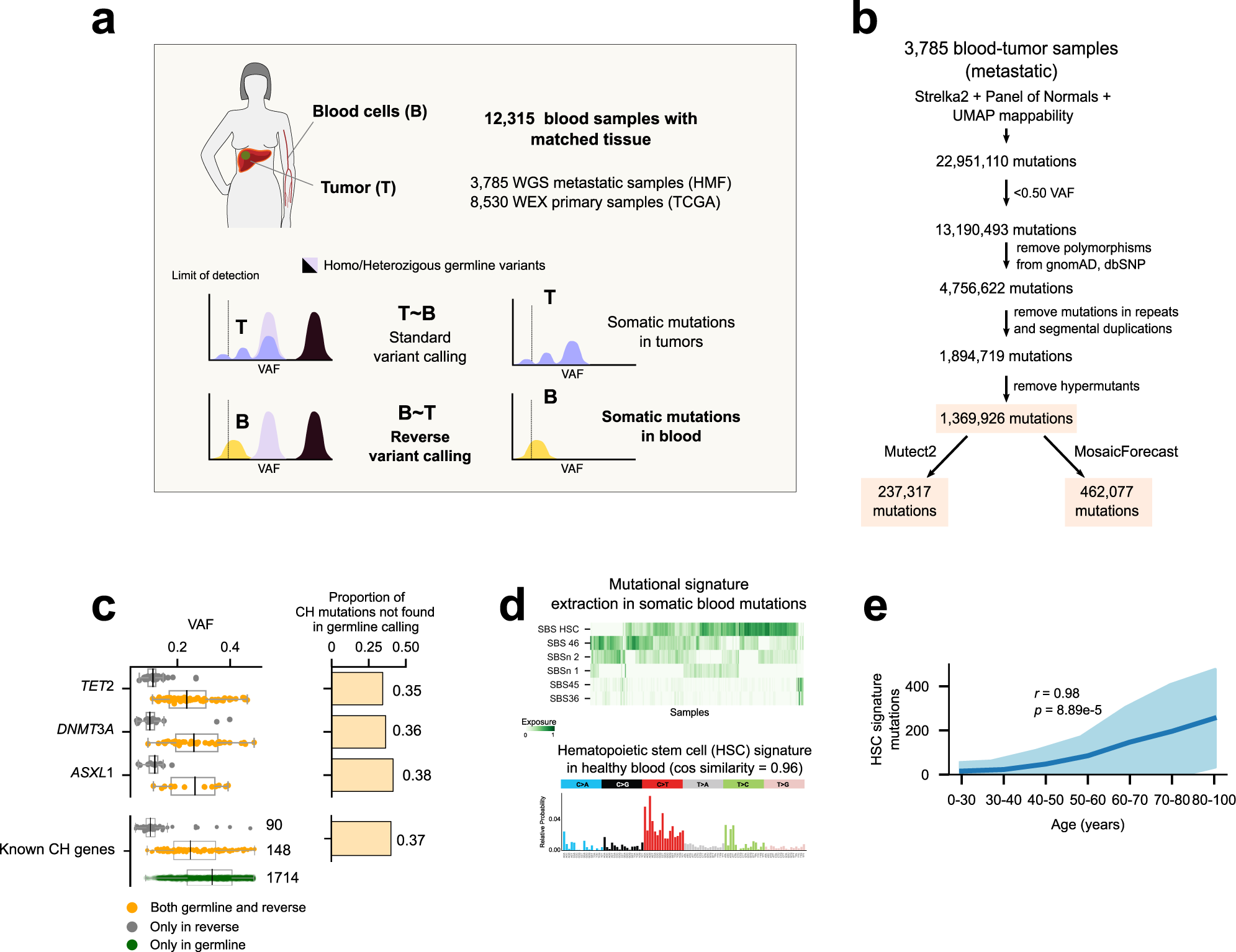
Discovering the drivers of clonal hematopoiesis

Whole genome error-corrected sequencing for sensitive circulating
Recomendado para você
-
![SCP 7143 J THE KNOB [SFW?]](https://i.ytimg.com/vi/hvxBDyQvZmc/maxresdefault.jpg) SCP 7143 J THE KNOB [SFW?]12 abril 2025
SCP 7143 J THE KNOB [SFW?]12 abril 2025 -
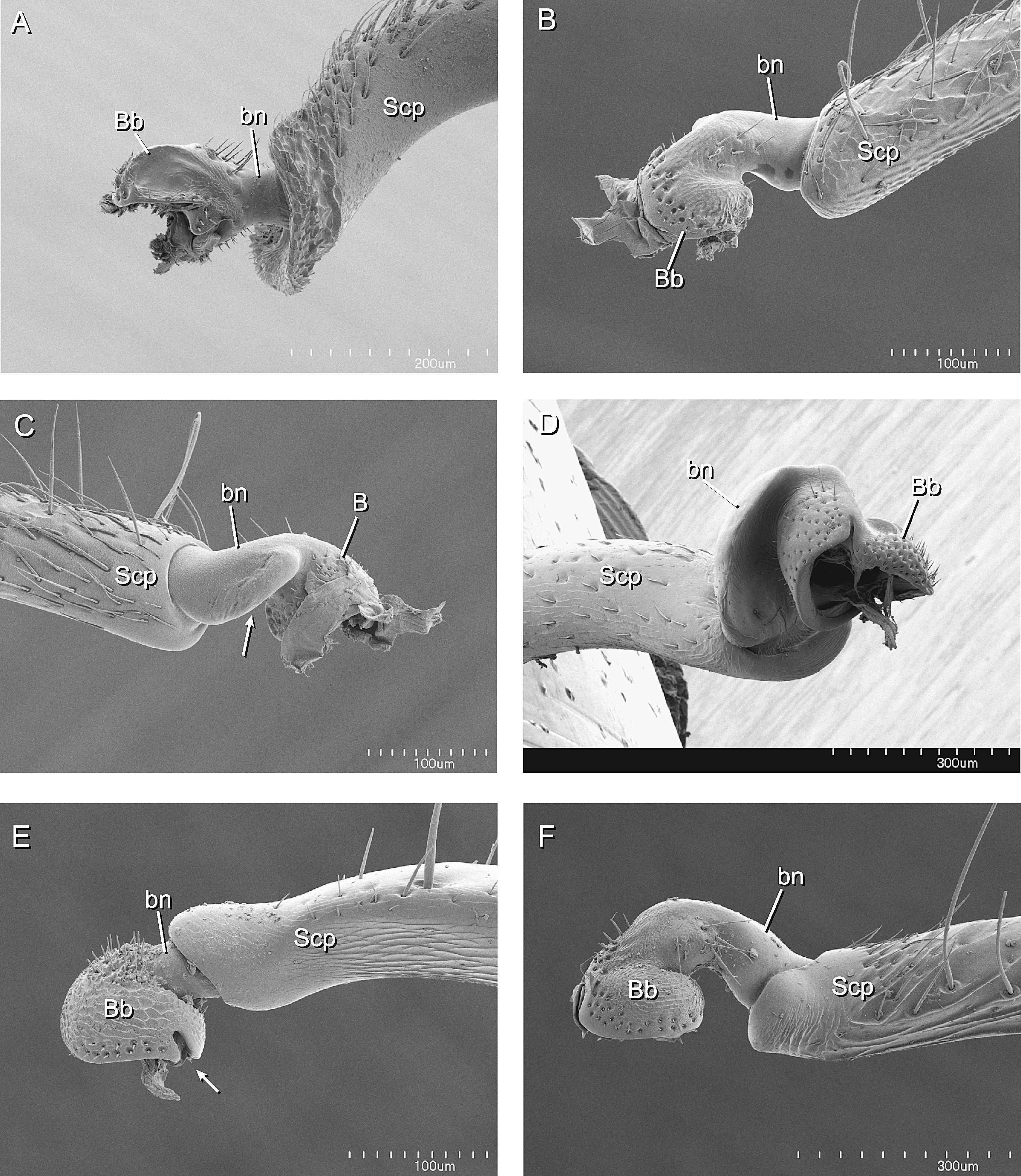 A Phylogenetic Analysis of Ant Morphology (Hymenoptera: Formicidae) with Special Reference to the Poneromorph Subfamilies12 abril 2025
A Phylogenetic Analysis of Ant Morphology (Hymenoptera: Formicidae) with Special Reference to the Poneromorph Subfamilies12 abril 2025 -
 Why my pp hard : r/DankMemesFromSite1912 abril 2025
Why my pp hard : r/DankMemesFromSite1912 abril 2025 -
 PDF) Tractographical model of the cortico-basal ganglia and corticothalamic connections: Improving Our Understanding of Deep Brain Stimulation12 abril 2025
PDF) Tractographical model of the cortico-basal ganglia and corticothalamic connections: Improving Our Understanding of Deep Brain Stimulation12 abril 2025 -
 SCP-175 Minecraft Skin12 abril 2025
SCP-175 Minecraft Skin12 abril 2025 -
parse_in/data.json at master · squiidz/parse_in · GitHub12 abril 2025
-
 Inhibition of HIV-1 Replication and Activation of RNase L by Phosphorothioate/Phosphodiester 2′,5′-Oligoadenylate Derivatives - ScienceDirect12 abril 2025
Inhibition of HIV-1 Replication and Activation of RNase L by Phosphorothioate/Phosphodiester 2′,5′-Oligoadenylate Derivatives - ScienceDirect12 abril 2025 -
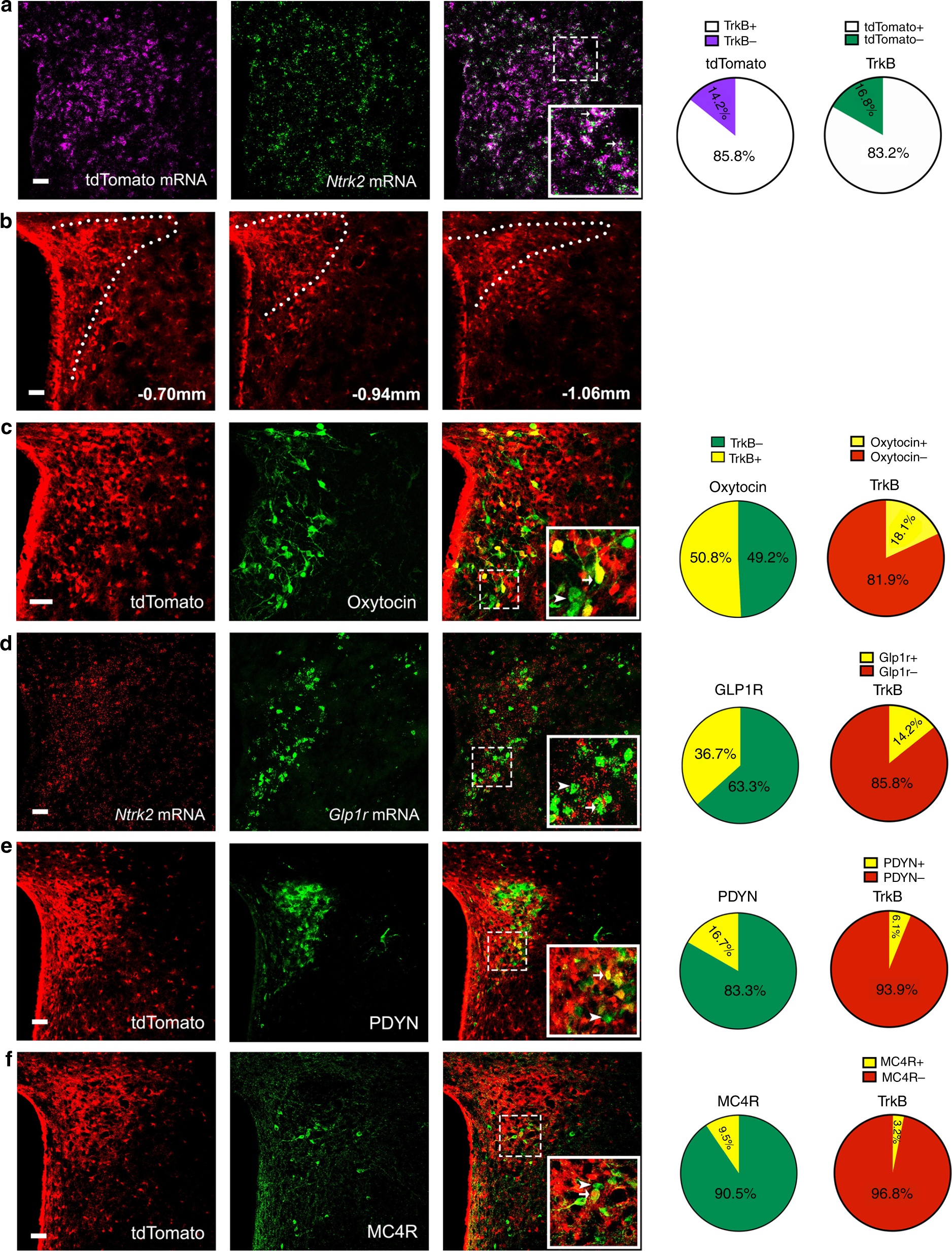 TrkB-expressing paraventricular hypothalamic neurons suppress appetite through multiple neurocircuits12 abril 2025
TrkB-expressing paraventricular hypothalamic neurons suppress appetite through multiple neurocircuits12 abril 2025 -
 Solubility Enhancement of Hydrophobic Substances in Water/Cyrene Mixtures: A Computational Study12 abril 2025
Solubility Enhancement of Hydrophobic Substances in Water/Cyrene Mixtures: A Computational Study12 abril 2025 -
 Journal of Comparative Neurology Systems Neuroscience Journal12 abril 2025
Journal of Comparative Neurology Systems Neuroscience Journal12 abril 2025
você pode gostar
-
 The Sad Reason We'll Never See A Real Pokemon MMO12 abril 2025
The Sad Reason We'll Never See A Real Pokemon MMO12 abril 2025 -
 O Phrasal Verb TO GIVE UP em inglês12 abril 2025
O Phrasal Verb TO GIVE UP em inglês12 abril 2025 -
 Microsoft Leak Included Roadmap For Unannounced Bethesda Games Including An Oblivion Remaster, A Fallout 3 Remaster, Dishonored 3, & More - mxdwn Games12 abril 2025
Microsoft Leak Included Roadmap For Unannounced Bethesda Games Including An Oblivion Remaster, A Fallout 3 Remaster, Dishonored 3, & More - mxdwn Games12 abril 2025 -
 Marvel Announces New Scarlet Witch Run Amid Solo Movie Rumors12 abril 2025
Marvel Announces New Scarlet Witch Run Amid Solo Movie Rumors12 abril 2025 -
 The Doors (1991) - IMDb12 abril 2025
The Doors (1991) - IMDb12 abril 2025 -
 Why Alan Wake 2 Isn't On Steam? - Gamer Tweak12 abril 2025
Why Alan Wake 2 Isn't On Steam? - Gamer Tweak12 abril 2025 -
 Arsenal fortitude pleases keeper Szczęsny, UEFA Champions League12 abril 2025
Arsenal fortitude pleases keeper Szczęsny, UEFA Champions League12 abril 2025 -
 Calma Coração Official Resso - Railena Show-Junior Vianna12 abril 2025
Calma Coração Official Resso - Railena Show-Junior Vianna12 abril 2025 -
 Todos los animes de Crunchyroll doblados al castellano12 abril 2025
Todos los animes de Crunchyroll doblados al castellano12 abril 2025 -
What is the meaning of Joga muito? - Question about Portuguese (Brazil)12 abril 2025